New technique improves detection of cancer DNA in blood

A team led by researchers at Broad Institute of MIT and Harvard, Dana-Farber Cancer Institute, and Harvard Medical School has developed a new method to identify thousands of DNA mutations accurately and efficiently in a patient's blood sample with minimal sequencing. The approach, called MAESTRO, could one day enable the detection of residual cancer in patients who have undergone treatment, alerting doctors to disease recurrence earlier and more cheaply than current techniques allow.
"The ability to findrare mutationsin a clinical sample is useful in many areas of biomedicine and diagnostics," explained co-senior author Viktor Adalsteinsson, associate director of the Gerstner Center for Cancer Diagnostics at the Broad Institute. "Current techniques require a great deal of sequencing to find low-abundance DNA fragments, whereas MAESTRO is sensitive enough to find thousands of mutations with a hundred times less sequencing."
这项工作发表在本周出版的Nature Biomedical Engineeringwith additional co-senior authors Todd Golub, director of the Broad Institute and faculty at Dana-Farber Cancer Institute (DFCI) and Harvard Medical School (HMS), and Michael Makrigiorgos, professor at DFCI and HMS.
Hunting for cancer DNA
In a patient with cancer,tumor cellsshed bits of their DNA into the blood—fragments with telltale mutations indicating they came from the diseased tissue. But any tumor DNA in ablood sampleis a tiny fraction floating in a sea of healthy genetic material. Accurately detecting this sparse amount of DNA is a challenge, especially when hunting for the small number of tumor cells left behind after cancer treatment (called "minimal residual disease," or MRD).
"We see this need for higher sensitivity," said Adalsteinsson. "And our team has been working on a number of technologies to address that."
The team has previously demonstrated success in detecting small amounts of residual cancer DNA fromblood samplesby scanning for hundreds of cancer mutations. Scanning for thousands of mutations can improve MRD detection rates even more, but this typically requires an enormous amount of sequencing to deliver accurate results. MAESTRO, which stands for "minor allele enriched sequencing through recognition oligonucleotides," is a more efficient approach to detect low-frequency mutations.
How MAESTRO works
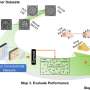
To use MAESTRO, researchers first sequence a patient's tumor biopsy to understand the landscape of mutations. With this information in hand, they can create specialized molecular probes that will bind to only those tumor-associated sequences of DNA. Scientists add the molecular probes to the cell-free DNA from blood samples, then wash away any unbound DNA, enabling the sequencing machines to pick out the rare cancer mutations from the sample.
In this study, MAESTRO performed just as well as more conventional sequencing approaches at detecting hundreds of low-abundance mutations, uncovering the majority while requiring significantly fewer resources. Additionally, MAESTRO enabled the team to increase the search to 10,000 mutations at low cost, profoundly boosting the detection results.
The researchers also reexamined patient samples that had been analyzed using their earlier methods. With MAESTRO, they uncovered substantially moremutationsfrom each tested blood sample, enhancing the detection of MRD aftercancer treatment.
"MAESTRO combines the advantages of depth and breadth in a single protocol," said Makrigiorgos. "This opens up the possibility of detecting MRD earlier, or identifying circulating DNA from cancers that shed very little."
"This project has been a great reminder that new methods can make DNA sequencing even more powerful," added Golub. "It will be exciting to see how MAESTRO can impact basic discovery and, in the future, clinical care."
Going forward, theteamis continuing to build a suite of technologies that can reduce the cost and increase the sensitivity ofcancerdetection, so that patients who may need additional treatment to prevent recurrence can be identified sooner.
更多的信息:Gregory Gydush et al, Massively parallel enrichment of low-frequency alleles enables duplex sequencing at low depth,Nature Biomedical Engineering(2022).DOI: 10.1038/s41551-022-00855-9